A DNA illustration from Anna Hupalowska
 A team from Columbia University present a microfluidic device that can capture single-cell lysates in picotire plates and produce single-cell 3' tag RNA-Seq library prep at $0.10 cents per cell: Scalable microfluidics for single cell RNA printing and sequencing. They discuss two methods - RNA “printing” on glass and RNA capture on beads, but the paper focuses on the second of these. This is an early paper with two experiments each of which could be improved by another round or two in the lab. However in this fast moving field I can’t help but wonder if this is a method that will never be commercialised and if the authors simply want to make their mark? I think the fact that single-cell methods are appearing thick and fast shows how much this space there still is to innovate into. And for the investment pundits looking to Fluidigm et al, there is a danger that a clever molecular biology approach could allow single cell without any hardware - imagine library prep inside a cell? How does it work: In the RNA capture on beads method cells are loaded into microwells, then barcoded oligo-dT coated beads are loaded such that one fills each picotitre well. cDNA synthesis on these beads generates a barcoded cDNA ready for downstream RNA-seq library prep. In the paper they provide proof-of-concept data on a five lane chip with >10,000 microwells, cells are loaded at low concentration and Poisson statistics ensure that >95% of wells with have 1 or 0 cells. Wells are sized such that only a single bead can also be accommodated. The chip is hydrated and washed with PBS, cells were loaded and settle by gravity, excess cells are removed by further PBS washes. The microwells are “closed” by oil and freeze-thaw was used to lyse cells releasing mRNA onto the beads. After mRNA capture all subsequent steps are performed with the chip “open”. They used the Cel-seq protocol where 1st and 2nd strand cDNA synthesis are followed by IVT. The resulting aRNA is collected and is used for a second RT using random hexamers, and this is followed by PCR to produce barcoded RNA-seq libraries for sequencing. Phew! Barcode production: They employ a very clever combinatorial barcoding method that allows just 106 oligos to produce 960 barcodes. First, beads are coated with oligos containing a T7 promoter and Illumina adaptor. Second, the beads are plated to allow 96 individual barcode adaptors to be annealed to these covalently attached oligos, and primer extension creates the full-length covalently attached 1st barcode oligo. Third, beads are pooled for a final step similar to round two but this a second round of primer extension using one of ten secondary barcoding oligo creates a final pool of 960 uniquely barcoded beads. Phew again!! from CoreGenomics blog  You'll probably remember the last time you had the flu, but what about that time you had measles – or was it chicken pox? Your blood knows: it keeps a record of every virus you've ever been infected with. A tiny drop of the stuff can now be tested to reveal a person's viral history. The test, called VirScan, reveals that adults around the world tend to have been infected by an average of 10 viruses over their lifetime. It could also be used to identify links between viral infections and mysterious diseases like chronic fatigue syndrome. When a virus infects us, our immune cells respond by producing antibodies that neutralise it when they bind to specific proteins on its surface. These antibodies continue to be made long after the virus has been cleared from our body, ready to mount a quicker response should it return. This means that their presence can act as a viral footprint – a clue that the viruses they target were once in the bloodstream. To test whether someone has been infected with a virus, expose a sample of their blood to a viral protein. If antibodies target it then the virus has infected the person in the past. Stephen Elledge at Harvard University and his colleagues have pushed this idea further and developed a way to test a blood sample for every single family of human virus in one go. Gotta catch 'em all Elledge says that all he needs to carry out the test is a tiny amount of a person's blood – less than a drop. It costs just $25, and could help doctors identify hidden infections. "A lot of people have hepatitis C, for example, without realising," says Elledge. You could imagine routinely screening people in this way, he says. To develop VirScan, Elledge and his colleagues used an international database to look up all viruses known to infect humans – around 1000 strains from 206 viral species. Using this information, they recreated the DNA in each virus that's responsible for making its proteins, and put the DNA segments into individual bacteriophages – viruses that infect bacteria. Each bacteriophage then manufactures a particular viral protein on its surface. When someone's blood is mixed with the bacteriophage brew, any circulating antibodies latch on to the associated proteins on the bacteriophages. Sequencing these bacteriophages then reveals the person's viral history. David Matthews at Bristol University in the UK thinks the best use of VirScan might lie outside of diagnostics, considering we already have quick and easy tests for individual viruses. "Usually when you've got a set of symptoms, doctors have a pretty good idea of what you've got," he says. Moreover, the immune system takes a while to make antibodies, so you might not find a strong antibody response in the early stages of an infection. The test would also not be able to distinguish between antibodies made as a result of an infection and those triggered by a vaccine. Disease detective Instead, the technique might be useful in outbreaks of new viruses. Understanding how our immune system responds to other viral fragments might reveal clues as to which family the new virus belongs to, says Pamela Vallely at the University of Manchester, UK. "If we'd have had this test during the HIV outbreak in the 1980s, it would have given us a clue for where to be looking to find out more about the virus," she says. "It's a really exciting technique." As well as playing an investigative role in outbreaks, VirScan could also offer a way to investigate whether viruses are involved in disorders that aren't well understood. For example, Elledge's team will be collaborating with another group to test people with chronic fatigue syndrome, to see if they might have been infected with any of the same viruses. "Multiple sclerosis is usually wheeled out as being linked to a virus," adds Vallely. "You could check." Down on the farm The team used the test to screen blood samples from 569 people from four countries – the US, South Africa, Thailand and Peru. As you might expect, adults appeared to have encountered more viruses than children. Each person had been infected with an average of 10 viruses over their lifetime. Matthews thinks it would be worth extending the screen to animal populations. He envisages screening wild populations of animals thought to be linked to emerging diseases. "You could test the wild bat population to get a good idea of what viruses are out there," he says. At the same time, farm animals could be comprehensively screened. Farmers that are able to identify viruses affecting the health or yields of their herds might be able to halt the spread of those viruses, says Matthews. "It is a fantastic piece of work and will be very, very useful," he says. Journal reference: Science, DOI: 10.1126/science.aaa0698 Newscientist MERS has come to South Korea in a big way. Cases more than doubled between 4 and 9 June, from 41 to 95, making it the second most infected country after Saudi Arabia, its epicentre. Seven people have died so far, but we know from outbreaks elsewhere that the virus is lethal in around 37 per cent of cases.
So far MERS remains a hospital-transmitted infection: nasty and expensive to treat, but in theory manageable. On Tuesday, the South Korean health ministry announced that it expects the outbreak to start declining this week. And a genetic analysis shows MERS is showing no signs of going pandemic – yet. That doesn't mean it isn't seeing the world. Since the virus emerged in 2012, ithas invaded 25 countries. But the vast majority of the 1190 known cases have been in Saudi Arabia and its neighbours. The first South Korean case was in a man who had visited Saudi Arabia, the UAE, Bahrain and Qatar. Third Wave South Korea is the first country outside the Middle East with more than a few cases, or where the virus has transmitted between three people in succession. The spike in cases this week is that third wave of people, who were exposed a week or more ago. But we are still only dealing with one chain of transmission, says Peter Ben Embarek of the World Health Organization. Every Korean infection since the first was caught in a hospital that had handled previous cases. The emergency department of Seoul's Samsung Hospital alone accounted for 10 cases last week. "The early symptoms are very non-specific," says Ben Embarek, meaning someone may not be diagnosed, and isolated, until they are quite ill. But the virus hasn't spread outside of hospitals, so once recognised it can, in theory, be contained by better hospital hygiene, and by isolating anyone the patient has been in contact with. Korean hospitals have been using CCTV footage to trace these people; nearly 3000 are currently in quarantine. The new SARS? Fears that MERS could go pandemic aren't unreasonable. It is a member of the coronavirus family and seems to have originated in bats – just like SARS, a virus that emerged in China in 2003 and spread worldwide, killing nearly 8500 people. But SARS behaved differently. Like MERS, it mainly infected hospital staff, but it could also spread in the general population. It invaded Hong Kong when a Chinese doctor passed it on to strangers in his hotel, and it later infected an apartment block via the ventilation system. MERS seems harder to catch. But hospitals are vulnerable: staff move from bed to bed; sick people have lower defences; medical procedures, such as removing breathing tubes, can spray droplets containing the virus, which is otherwise trapped deep in the lungs. Some infections, like the antibiotic-resistant bacteria MRSA, specialise in hospitals. MERS could fill a similar niche if it is accidentally released in more hospitals around the world. In countries less wealthy than South Korea, it could be a struggle to dislodge it. More like MRSA That may be less scary than a MERS pandemic, but MRSA shows why it would still be bad news. It is half as lethal as MERS, and unlike MERS can be killed by some drugs. Yet it still costs European hospitals €380 million a year, while killing some 300 people annually in England and Wales alone. Virologists say MERS won't go pandemic unless it becomes better at transmitting between people. So far the virus is showing no sign of evolving that capability. In a genetic analysis published this week, virologists found the Korean virus is virtually identical to Saudi strains, suggesting we can breathe easy on that front for now. Newscientist Who you are is not just down to your DNA; your environment plays a big role, too. Lifestyle factors such as stress and diet can alter the way your genes are tagged with on and off switches, which modify the way your genes are expressed. While this much was known, how these changes seem to be passed on to future generations has puzzled scientists. Now, a new study has finally provided some insight into what’s going on.
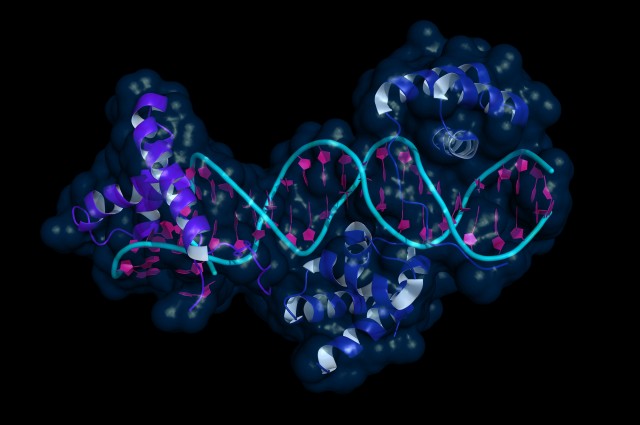
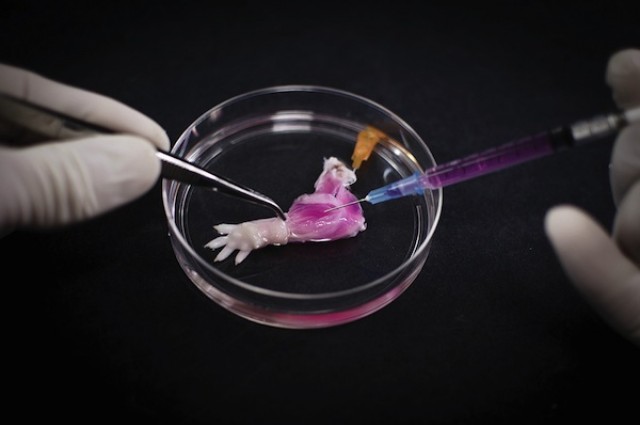
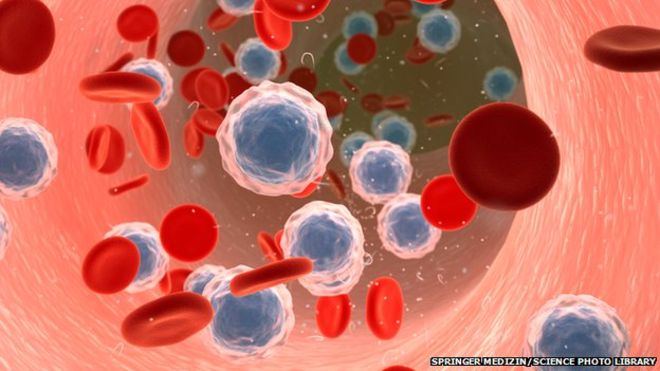
Although early cells destined to become eggs and sperm are wiped of these changes early on in embryo development, scientists have revealed that some stretches of DNA resist this so-called reprogramming, allowing the modifications to persist and thus become heritable. Importantly, the researchers discovered that some of the resistant genes are associated with certain diseases, including obesity and schizophrenia. These intriguing findings have been published in the journal Cell. While DNA contains the codes necessary to build an organism, not all of our genes need to be active at the same time or in the same place throughout the body. This is where epigenetics come in; these modifications to DNA can alter which genes are turned on or off without changing the actual DNA sequence itself. For example, a chemical tag called a methyl group can be added or removed that can inactivate genes by making the DNAinaccessible to the machinery that comes along to read it. This process of DNA methylation continues throughout our lives, but it can also occur in response to factors in the environment. For example, stress such as famine can alter methylation patterns, and an increase in the risk of schizophrenia has even been found in girls born to mothers who experienced prolonged periods of famine during pregnancy. But it doesn’t end there; mice subjected to stress in the lab have been shown to produce two generations of depressed offspring, even though their upbringing was not stressful. These observations are perplexing because epigenetic data was thought to be erased in the cells that give rise to sperm and egg cells, called germ cells, to prevent any abnormal methylation from accumulating that could harm offspring. In an attempt to solve this apparent mystery, scientists from the University of Cambridge began examining the processes at play in developing mice embryos. In particular, they were interested in the embryo’s germ cells since these are what will eventually give rise to the animal’s offspring. They found that the process of germ cell reprogramming takes place over a period of roughly seven weeks, kicking off around two weeks into embryo development. This clearance phase involves the initiation of an inhibitory network that prevents the action of enzymes that facilitate epigenetic changes or their maintenance. However, they found that around 5% of the genome resisted reprogramming. This means that any methylation that has occurred in these regions is not removed and can thus persist, potentially impacting future generations. Upon closer inspection, the researchers revealed that some of these resistant regions are associated with certain conditions, including diabetes, obesity and schizophrenia. These reprogramming “escapees” could therefore help explain why environmental factors can not only impact an individual’s health, but the health of their future generations. http://www.iflscience.com/  In the latest step towards transplantable bioengineered parts, researchers have built rat forelimb tissue – complete with working blood vessels and muscle fibers – in the lab. After they transplanted the biolimb into a recipient rat, the blood vessels filled with circulating blood, and the muscles even flexed the rat’s wrists and the joints in its paws. For people who have lost a limb, transplants could help to improve the quality of life. But this also means having to take immunosuppressant drugs so that their bodies don’t attack the donated parts. That’s why a lot research has focused on using the patients’ own stem cells to regenerate their own replacement tissues, but what’s been missing so far is the scaffold (or matrix) to provide shape and support for growing cells as they become the complex tissues that make up a limb. So, a team led by Massachusetts General Hospital’s Harald Ott tried stripping away cells from an existing rat forelimb and then repopulating the remaining matrix with progenitor cells. This decellularization technique has previously been used to build bioartificial organs like kidneys, livers, hearts, and lungs in animals, but engineering tissues for a bioartifical limb is a different kind of task. “The composite nature of our limbs makes building a functional biological replacement particularly challenging,” Ott explains in a news release. “Limbs contain muscles, bone, cartilage, blood vessels, tendons, ligaments, and nerves – each of which has to be rebuilt and requires a specific supporting structure called the matrix.” First, they used a detergent to strip all the cellular material from forelimbs taken off deceased rats, but they made sure to keep the primary vasculature and nerve matrix. They spent a week removing all the cellular debris. Meanwhile, muscle and blood vessel progenitor cells were cultured separately. When the forelimb matrix was cell-free, the team suspended it in a bioreactor that provides a nutrient solution and electrical stimulation to promote growth (pictured to the right). Vascular cells were injected into the limb’s main artery to regenerate veins and arteries, while muscle cells were injected directly into sheaths within the matrix that help define each muscle (pictured above). Two to three weeks later, when the forelimb was taken out of the bioreactor, the team looked for the presence of vascular cells along the blood vessel walls and muscle cells properly aligned throughout the muscle matrix. To see if the new limb was functional, they applied electrical stimulation to the muscle fibers, which contracted with a strength that’s 80 percent of what you’d expect from a newborn rat. After the limb was transplanted into recipient rats, the vascular system quickly filled with blood that continued to circulate. With a bit of stimulation, the muscles within the graft flexed the rats’ wrists and the digital joints of the rats’ paws. Next up, regrowing nerves! The work appears in the latest issue of Biomaterials. photo credit: A suspension of muscle progenitor cells is injected into the cell-free matrix of a decellularized rat limb. Bernhard Jank, MD / Ott Laboratory, Massachusetts General Hospital Center for Regenerative Medicine from iflscience.com Scientists say they have found evidence that cancer cells can go to 'sleep', avoiding the effects of chemotherapy, and then 'reawaken' years later.
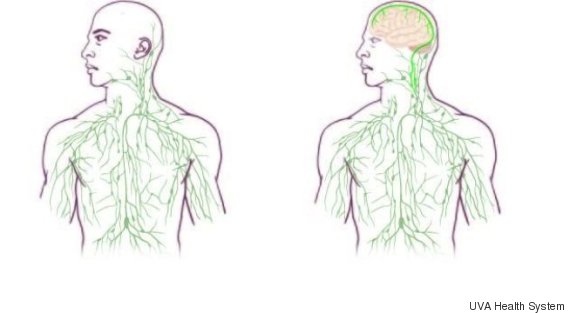
Researchers at the Institute of Cancer Research say this may explain why some cancers return, many years after they appear to have been cured. They analysed a patient whose leukaemia returned after 20 years in remission. The findings may help scientists to root out these dormant cancer cells, wake them up and kill them. The study, published in the journal Leukemia, found that the cancer cells which 'woke up' in the patient after a period of two decades were similar to a group of cancer cells that pre-dated the original bout of the disease. Blood and bone marrow samples were taken from the patient when he was diagnosed with a rare form of leukaemia at four years old and compared to samples taken when he relapsed aged 25. In future it might be possible to speed up the growth of these pre-cancerous dormant cells so that they can be targeted and killed.Prof Mel Greaves, The Institute of Cancer Research Researchers identified a specific DNA mutation in cancer cells from both blood samples, in which two genes called BCR and ABL1 fuse together. They said this showed a common link between the original and the relapsing leukaemia. But they also found many new genetic changes had occurred in the cancer cells when the patient relapsed. This implies that cancer cells had become dormant, resisted chemotherapy and then 'woke up' after many years of rest. The cells may have survived because they were growing much more slowly than other cancer cells - and chemotherapy attacks rapidly dividing cells. Study leader Professor Mel Greaves, director of the Centre for Evolution and Cancer at The Institute of Cancer Research in London, said the research showed that cancer cells are cunning. "It provides striking evidence of cancer evolution in action, with cancer cells able to lie dormant to avoid treatment, and then to accumulate new mutations capable of driving a new bout of disease. "Blood stem cells regularly fluctuate between being dormant or 'asleep' and dividing very quickly, so it seems cancer cells are just borrowing this trick to avoid being killed by chemotherapy." Prof Greaves added: "In future it might be possible to speed up the growth of these pre-cancerous dormant cells so that they can be targeted and killed using chemotherapy, to reduce the risk of relapse even further." Dr Matt Kaiser, head of research at Leukaemia and Lymphoma Research, said there were still too many children whose cancer returns. "If we can build up a picture of what causes rare cases of late relapse and how we can detect and prevent it, we may be able to deliver more true cures for this terrible disease." BBC news Neuroscientists have uncovered a previously unknown direct connection between the brain and the immune system -- a finding that could have significant implications for the treatment of brain disorders like Alzheimer's disease, multiple sclerosis and autism.
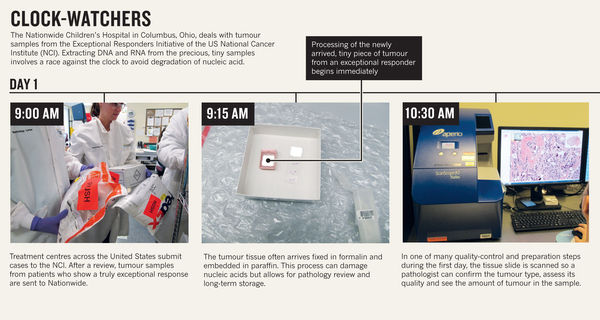
The discovery came as a surprise to Dr. Kevin Lee, chairman of the University of Virginia's neuroscience department. “The first time these guys showed me the basic result, I just said one sentence: ‘They’ll have to change the textbooks,'” Lee said in a press release Monday. The study's lead author, Dr. Jonathan Kipnis of the University of Virginia's Center for Brain Immunology and Glia, echoed the sentiment. "When we discovered the lymphatic vessels we were very very surprised, because based on the textbooks -- these vessels do not exist," Kipnis said in an email to The Huffington Post. A direct connection. While previous research held that there was no direct connection between the brain and the lymphatic system, the new findings, which were recently published in the journal Nature, present a model of the lymphatic system that includes the brain. Though not a part of the immune system, the lymphatic system carries lymph, a clear fluid filled with white blood cells that helps remove toxins from the body. The lymphatic system is connected to every other system in the body, and was believed to end at the base of the skull. But the researchers noticed something strange while looking through their microscopes at slides containing the membranes of mice's brains. They saw lymphatic vessels, which had never before been observed above the base of the skull. Why have they never been observed before? Kipnis explained that the vessels are "well hidden" behind a major blood vessel to the sinuses in an area of the brain that's difficult for scientists to image. While the observation was made in mice, the same anatomy is believed to exist in humans as well. New questions. A next step of the research is to determine how the vessels might be involved in diseases involving the brain and the immune system, such as multiple sclerosis and Alzheimer's. "We believe that for every neurological disease that has an immune component to it, these vessels may play a major role," Kipnis said. Though the findings are preliminary, the researchers hope they'll open up a number of new possibilities for treating these and other neurological disorders through therapies that target the lymphatic vessels of the brain. For example, Kipnis explained that the findings could shed light on why large protein chunks accumulate in the brains of Alzheimer's patients. "We think they may be accumulating in the brain because they're not being efficiently removed by these vessels," he said. Other members of the scientific community have expressed excitement about the discovery, while cautioning that the findings have not yet been replicated or confirmed. "With further research, the findings could help scientists to target inflammation in the brain, which is a factor in many neurological conditions," James Nicoll, professor of neuropathology at the University of Southampton in England, told The Guardian. Huffingtonpost.com Nature 520, 389–393 (16 April 2015) doi:10.1038/520389aPublished online 15 April 2015 Sometimes a drug causes a tumour to completely recede, but only in a tiny percentage of people. Scientists want to decipher such outlier responses for the benefit of all patients. If Patient X were like most people with advanced bladder cancer, she would probably be dead by now. After her first diagnosis, she received standard chemotherapy. It failed. Then she entered a clinical trial for a drug that was originally approved to treat other tumour types: would it also work in metastatic bladder cancer? Apparently not — none of the other patients in the trial did well. Yet Patient X thrived. Her tumour completely disappeared, says computational biologist Barry Taylor at Memorial Sloan Kettering Cancer Center (MSKCC) in New York, where Patient X was treated. Today, a little more than five years after treatment, she is healthy and has no evidence of disease1. Patient X (her identity is shielded to protect her privacy) is an exceptional responder, one of those rare individuals who have a dramatically positive response to a therapy that does little or nothing for most other patients. This response is not unique to cancer. Immunologists, for example, have discovered why some individuals can be HIV-positive and yet avoid the symptoms of AIDS. By definition, exceptional responses are rare, which makes them hard to study. Their anecdotal nature seems to contradict the teachings on statistically sound results in biomedical research. In a clinical trial, even if there are several exceptional responders, a drug will fail to achieve approval because it does not improve the health of the majority of patients. This means there has been little incentive for researchers or drug companies to investigate thoroughly why a few people respond so well. “The hope is that a signal seen in an exceptional responder will be seen in other cancer patients.” But that neglect is starting to be addressed as more cases of exceptional responses in cancer reach the published scientific literature and techniques emerge for profiling patients at the molecular level2. In Patient X's case, genome sequencing revealed a mutation in her tumour that explains why her cancer is specifically vulnerable to the drug she received on the clinical trial1. Such successes indicate that searching for and profiling these patients can potentially help researchers to predict many other patients' responses to potential therapies. The relatively new ability to comprehensively characterize a tumour's genome, transcriptome (its gene expression) and metabolome (its metabolic processes) increases the chance of discovering the reasons behind outlier results, says Kenneth Kinzler, a cancer researcher at the Johns Hopkins Kimmel Cancer Center in Baltimore, Maryland. “The hope is that a signal seen in an exceptional responder will be seen in other cancer patients and be a predictor of therapeutic response regardless of tumour type,” he says. The exceptional profileThere is no universally accepted definition of exceptional responders, says Barbara Conley of the US National Cancer Institute (NCI) in Rockville, Maryland. Conley directs the Exceptional Responders Initiative (ERI), which profiles these patients. The ERI considers a drug response to be exceptional when a tumour disappears or when a patient shows an exceptional response to treatment and lives longer than 90% of others treated similarly. In tough-to-treat and advanced cancers, an exceptional response is when treatment causes a tumour to regress by at least 30%for at least six months, but only in less than 10% of people on the same treatment. In the case of Patient X, for example, her sequenced tumour genome revealed a mutation in a gene called tuberous sclerosis complex 1, which is one of several genes involved in a pathway that regulates cell growth and proliferation. The drug that worked for Patient X, but not for the other patients in the clinical trial, inhibits signalling in that pathway. But that does not completely explain Patient X's exceptional response. Analysis of tumour samples from 13 other patients in her trial showed that four had a mutation in the same gene, but the drug gave them only a short reprieve. To get a better understanding of Patient X and other exceptional responders, the ERI wants to do comprehensive profiling of a wide variety of parameters, including patients' clinical history, DNA changes, RNA levels of different genes (which reflect their activity) and metabolic pathways. Taylor and his colleagues have long encountered the critique that studying exceptional responders is merely generalizing anecdotes. But even though published studies on exceptional responders are few, he says, “I think the weight of evidence has now shifted that view.” Vincent Miller, a former MSKCC oncologist, agrees that views about outliers are changing and thinks that many more such individuals might be found. Any oncologist has a handful of patients in whom cancer just melts away with no obvious explanation, says Miller, who is chief medical officer of Foundation Medicine in Cambridge, Massachusetts, a company that performs genomic analysis of samples from people with cancer. In January, the pharmaceutical company Roche, based in Basel, Switzerland, bought a majority stake in Foundation Medicine, which is also involved in the ERI. The ERI encourages clinicians to get in touch if one of their patients has an exceptional reaction to a drug. At that point, a multidisciplinary review determines whether a more comprehensive profile is warranted, says Conley. In approved cases, and with the patient's consent, the physician sends in the complete medical record and a tumour sample. Around 160 submissions are currently under review. Conley and her team have been surprised to see submissions about established drugs as well as drugs still under development. The ERI makes sense only because large-scale sequencing efforts such as The Cancer Genome Atlas (TCGA) now offer huge data stores, says David Wheeler, who leads the ERI genome-analysis team at the human genome sequencing centre of Baylor College of Medicine in Houston, Texas. From Baylor, genome data will go to a database that is accessible by the research community. The first few ERI samples are now beginning to arrive at Baylor, and researchers there are all set to potentially perform whole-genome sequencing using their newly arrived equipment — HiSeq X Ten Illumina sequencers. Whole-genome sequencing is ideal, says Wheeler, because it provides the most complete genomic information. But it also requires enough sample and plenty of time and money; so when the samples are smaller or when only ones with lower tumour purity are available, the team will just focus on protein-coding genes, which make up the exome. For now, the ERI is in a pilot phase. If it proves successful, it could be scaled up by, for example, helping cancer treatment centres to forage for exceptional responders in their biobanks. But the pilot faces a few challenges. One key issue is time, says Kristen Leraas, who is the sample coordinator at the biospecimen processing facility of the Nationwide Children's Hospital in Columbus, Ohio, where all of the ERI's samples are processed (see 'Clock-watchers'). When a sample comes in, she says, scientists have to race against the clock to process, standardize and prepare it for sequencing: DNA and RNA have to be extracted from the sample quickly to avoid any kind of degradation. “We pretend our hair is on fire and we make sure we extract right away.” Another challenge is that exceptional responses are unexpected, so the cancer centres sending tissue samples to Columbus do not collect them in a standardized way. One sample might be blood from someone with leukaemia, whereas another might come from a solid tumour. And unlike the case with the TCGA, it might arrive without a matched healthy tissue sample from the same person. A sample might be smaller than a pencil eraser and, in some cases — when it comes from a fine-needle biopsy, for example — it might even be invisible to the naked eye. TCGA samples weigh on average 260 milligrams, whereas “if we get 100 milligrams, that's a lot”, says Jay Bowen, who directs logistics and data management at Nationwide's biospecimen processing facility. “Sometimes we make do with about 20 milligrams.” Comprehensive testingThe Nationwide laboratory's top priority with these samples is to extract enough nucleic acid to allow multiple analyses, including exome and messenger RNA sequencing. Some DNA is also sent to Foundation Medicine, where tests can detect and validate four classes of DNA alterations at once: substitutions of bases along the DNA strand, genetic insertions and deletions, changes in the number of copies of genes present in the genome and structural rearrangements3. Ideally, if the sample yields sufficient quantities of nucleic acid, whole-genome sequencing or other types of tests, such as analysis of DNA methylation, can be performed. A potential complication is that tumour samples taken during surgery or biopsy are often fixed in formalin and then embedded in paraffin. These formalin-fixed paraffin-embedded (FFPE) samples are standard in medical centres and are preferred by pathologists, who can easily shave off a thin slice when they want to study the tumour's cellular morphology under a microscope as part of diagnosis. But this process can crosslink nucleic acids, and can also oxidize and shear these molecules, says molecular biologist Erik Zmuda, who directs molecular characterization tasks at the Nationwide's biospecimen processing facility. There is a risk that a genomic signal indicative of an exceptional response is actually an FFPE artefact. Thus, for studying the tumour's genome, researchers much prefer frozen tissue. Zmuda and his colleagues at Nationwide and other institutions think they see a way to allow pathologists to continue to use their preferred FFPE preservation method while providing molecular biologists with the ability to profile a sample at the resolution they need. The team's idea is to find a telltale signature of FFPE artefacts in tumour samples, which would allow them to computationally mask these effects in the data. The team is developing an algorithm that would correct for the artefacts and thus make it easier to compare data from FFPE and frozen samples. That, in turn, could open up possibilities to retroactively analyse patient samples from pathology departments in any hospital. As well as helping the hunt for signals in outlier genomes, this method could also be adapted for use in genome analysis more generally when diagnosing and treating patients. Other fields have a longer tradition than cancer research does of looking at exceptional responders, says Stephen Friend, a former director of the oncology division of pharmaceutical company Merck in Kenilworth, New Jersey. Early in the AIDS epidemic, for example, immunologists noticed that some people can be HIV-positive but lack symptoms. This exceptional biology was found to result from a mutation that changes a protein on the surface of the immune-cell type that HIV infects, thus stopping HIV from entering the cell4. Such links between a specific mutation and disease have sometimes led to targeted drugs. But genomics is not a black and white world in which certain mutations lead to the same clinical course in all patients, says Friend. Environmental factors and other genetic variants play their part too. This may be why these targeted drugs do not work in 100% of the patients with that mutation, he says. Friend co-directs the Resilience Project (http://resilienceproject.me), which is geared towards finding outliers in many diseases5. The goal is to find people who harbour DNA changes that cause severe and rare childhood diseases, or that heighten cancer risk, but who have lived into healthy adulthood in spite of their genomes. The programme is run by the non-profit organization Sage Bionetworks, which is based in Seattle, Washington, and is devoted to setting up platforms through which scientists can collaborate and share data. The Resilience Project currently consists of researchers from the Icahn School of Medicine at Mount Sinai Hospital in New York (conversations are also under way with the Gurdon Institute in Cambridge, UK). DNA analysis is in progress on samples from more than half-a-million donors, says Friend, who also directs Sage Bionetworks. If the first analysis of the donor DNA reveals a mutation that could have killed the carrier, researchers can dig deeper into that person's genetics and biochemistry in an effort to understand their resilience. If one mutation is decisive, analysis can be quick, says Friend. But a mutation might act in conjunction with secondary mutations elsewhere in the genome. Searching for such mutation combinations is difficult, he says. But with an outlier genome in hand, researchers are at least trawling through a bucket of data, not an ocean of data. Scientists tend to keep findings under wraps until they publish. But Friend thinks that analysis should be a collaborative task that is spread across multiple laboratories. This would increase the speed at which scientists can decipher which factors — be they genetic, immunological, environmental or a combination — have protected resilient individuals. “What I'm hoping is that we can get scientists to take it on as a sort of crowd-sourced federated approach,” says Friend. “No one is paid to do that, no one owns the data.” Rare signalsIn a clinical trial, scientists strive for numbers: making sure there are sufficient cases of disease and controls to see whether a drug is having an effect, for example. They look for global trends rather than focus on the outliers, says Gustavo Stolovitzky, a researcher for the technology firm IBM in Yorktown Heights, New York, who runs the Dialogue for Reverse Engineering Assessment and Methods, a research venture and competition that, for example, looks at how well different algorithms predict the reaction of cancer cells to drugs6. By definition, outliers are too rare to have much statistical power, Stolovitzky says, and are usually dismissed as flukes. But conversely, he says, an exceptional response is a strong signal that is hard to miss. If many scientists hunt for exceptional responders in data from the ERI or the Resilience Project, perhaps 20 or even 50 cases can emerge. “That's starting to be something,” says Stolovitzky. “It's a number we can do statistics with.” If so, it may be possible to glimpse patterns that can help to explain how exceptional responders beat the odds. In profiling outliers, scientists will not know which of the molecular signals is decisive, which is why comprehensive profiles are needed for everything — genome sequencing data, gene expression data, clinical data and other assay results. Comparing these profiles is tricky: for example, it can often be a challenge to compare genomic sequence, says Trey Ideker, a computational biologist at the University of California, San Diego. “We sequence this individual and they're a snowflake,” he says — showing patterns that are unique even though the patients have the same type of cancer. Ideker says that one approach to address that diversity is to view cancer as a disease of pathways, in which groups of genes act together to perform functions in the cell. When analysed on a pathway level, he says, patterns do emerge. For example, researchers may find that dissimilar-seeming mutations in a cancer all fall in a certain pathway, meaning that they all impair the same cell function. These network patterns are not complete biochemical explanations of an exceptional response in cancer treatment, says Ideker, but they are indications of what to explore next. Crucially, he says, by considering pathways, an exceptional responder becomes part of a group. Even if it is not a large group, the person is no longer an outlier. Many patients could benefit from ventures to decipher the molecular profile of exceptional responders. A physician might realize that a drug that was not expected to do well in a given patient might actually be surprisingly suitable, says Taylor. This approach to cancer treatment complements an emerging idea that rather than focusing on the organ in which the tumour originated, treatments should be targeted to the molecular profile driving a given cancer. For research on outliers to be of greatest help, the outlier cases must be rigorously selected. Only then can the analysis deliver sound results despite the fact that it remains a profile of only one person, says Friend. Taylor agrees, pointing out that molecular analysis of tumours from patients is increasingly possible and that there is growing acceptance of studying outlier patients. “Nevertheless,” he says, “it requires that we stay focused on exploring the most significant outlier responses to ensure the greatest return for patients.”
Tiny treasure: The future of nano-gold Lumps of gold moulded into rings, coins and ingots have been highly prized for millennia. But recently, scientists have realised that tiny pieces of this precious metal – far too small to be seen by the naked eye - could also become a valued commodity. In labs around the world, gold nanoparticles are being tested as components in technology and medicines. See how gold could be used to kill cancer cells, improve the efficiency of solar cells and catalyse chemical reactions. Nature |
|||||||





 RSS Feed
RSS Feed